Overall goal
Our research is driven by the observation that the outcomes of lymphoma patients are extremely variable. Certain patients are seemingly cured, or enjoy very long remissions, while other patients experience disease relapse or refractoriness. By and large, we are currently unable to offer treatment that targets the therapeutic vulnerabilities that underlie a given patient’s lymphoma. We would like to change this. We postulate that the comprehensive deciphering of the biological underpinnings of lymphoma will lead to improved therapies for lymphoma patients.
In order to achieve this goal, we are applying state-of-the-art genomic techniques to primary patient samples to unravel tumour heterogeneity and to develop novel, innovative biomarkers to predict outcome in lymphoma. Furthermore, we are leveraging novel findings from discovery platforms to elucidate mechanisms of lymphoma pathogenesis, tumour evolution and treatment resistance.
Our ultimate goal is to improve patient outcomes through a better understanding of the diversity of responses to treatment and by tailoring therapy to each individual patient.
Research projects
1. Predicting outcome of follicular lymphoma using genomic approaches
Follicular lymphoma is clinically heterogeneous and treatment decision rely on clinical indices. However, the catalogue of genetic alterations observed in any given patient may provide the substrate for reponse following treatment. We mine existing and new datasets to identify novel genetic outcome predictors. The ultimate goal is to develop clinical-grade biomarkers that help us personalize therapies.
2. Understanding treatment resistance
Despite the consideration that treatment resistance ultimately contributes to lymphoma-related mortality, it is very poorly understood. Using patient-derived datasets, we try to identify genetic and phenotypic markers that are robustly associated with treatment resistance. Using functional genomics, we aim to understand how such markers cause resistance.
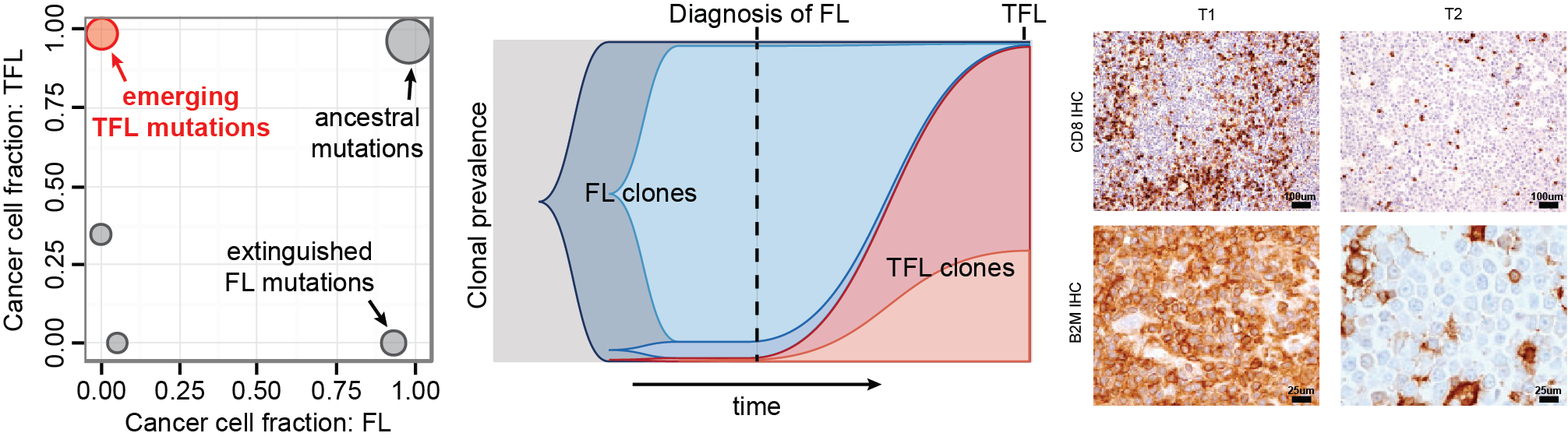
3. Deciphering tumour evolution
Clonal evolution is a fundamental property of how cancers evolve over time. We postulate that better understanding of clonal dynamics at play will provide us with new strategies to circumvent the emergence of resistant clones. We are trying to answer key questions related to intratumoral heterogeneity and tumour evolution using unique resources and datasets available at the Princess Margaret Cancer Centre.
4. Translational research platform
We aim to provide a comprehensive genomic research platform for clinical trials. This includes profiling of circulating tumour DNA, plasma biomarkers and tissue biopsies.